Publication
Variable selection for dimensionality reduction of biological datasets through bootstrapping of correlation networks
D.G. Aragones, M. Palomino, J. Sicilia, G. Crainiciuc, I. Ballesteros, F. Sánchez-Cabo, A. Hidalgo, G.F. Calvo
Computers in Biology and Medicine 168, 107827 (2024).
MOLAB authors
Abstract
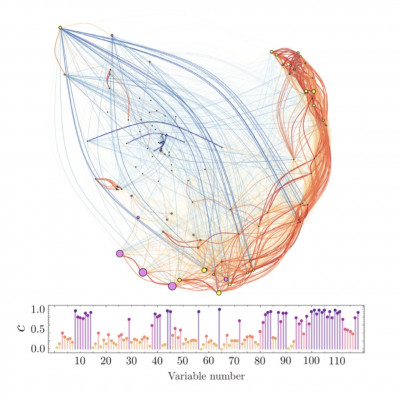
Identifying the most relevant variables or features in massive datasets for dimensionality reduction can lead to improved and more informative display, faster computation times, and more explainable models of complex systems. Despite significant advances and available algorithms, this task generally remains challenging, especially in unsupervised settings. In this work, we propose a method that constructs correlation networks using all intervening variables and then selects the most informative ones based on network bootstrapping. The method can be applied in both supervised and unsupervised scenarios. We demonstrate its functionality by applying Uniform Manifold Approximation and Projection for dimensionality reduction to several high-dimensional biological datasets, derived from 4D live imaging recordings of hundreds of morpho-kinetic variables, describing the dynamics of thousands of individual leukocytes at sites of prominent inflammation. We compare our method with other standard ones in the field, such as Principal Component Analysis and Elastic Net, showing that it outperforms them. The proposed method can be employed in a wide range of applications, encompassing data analysis and machine learning.